CRISPR/Cas: Programming the code of life just go exponentially easier
Origins
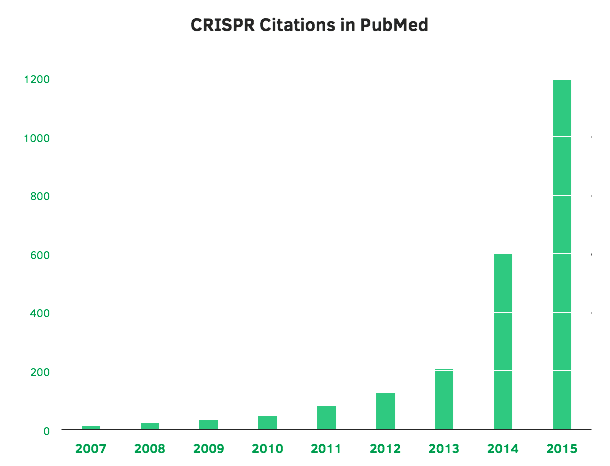
Since the discovery of heredity, scientists were looking for ways how to influence what traits were passed from one generation to the next. Although gene editing was possible in the past, the methods used so far were cumbersome, costly or error prone.
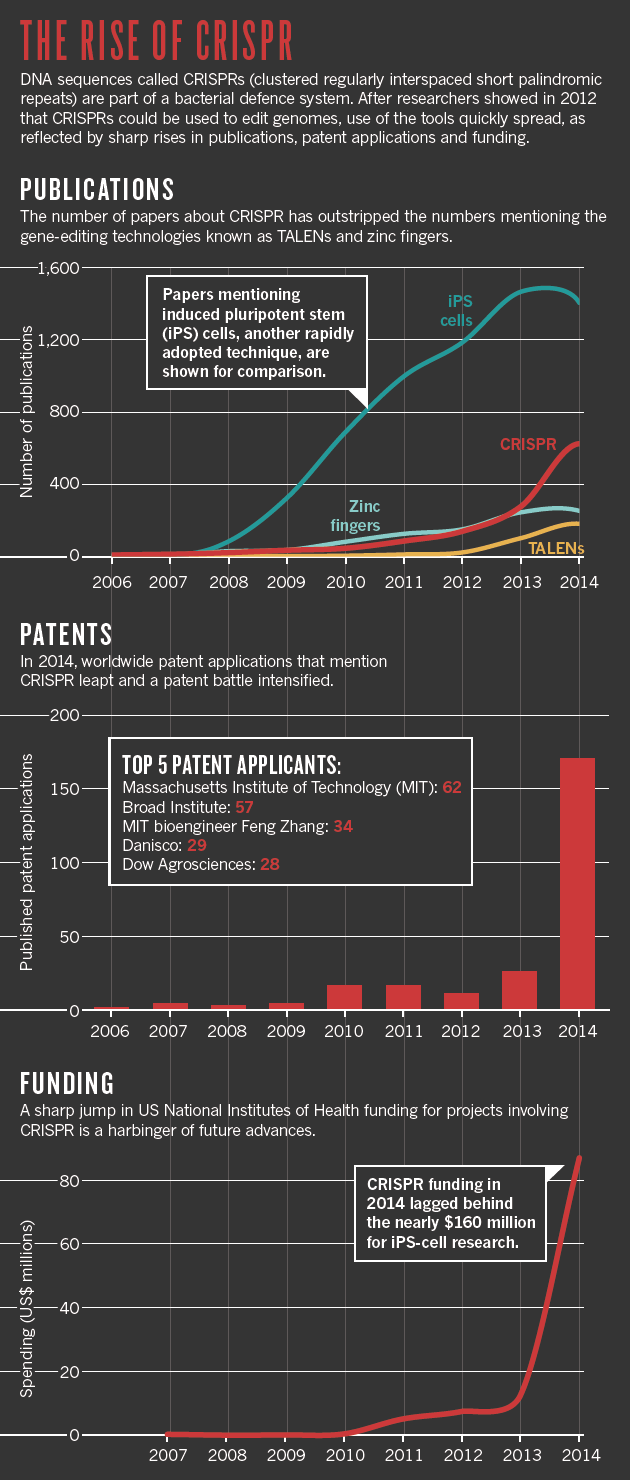
The discovery of a curious region in the genes of obscure bacteria over 20 years ago led to the development of the most convenient and precise instrument for gene editing so far: CRISPR. Since its first application in non-bacterial, or eukaryotic, cells, application of the technique increased exponentially.
This is part 3 of my series about exponential technologies in healthcare. In part 1 I am discussing why the convergence of several exponential technologies represents the largest opportunity of our time.
Part 2 is looking at what has enabled the exponential increase in genome data and what it means for healthcare.
CRISPR
The Clustered Regularly Interspaced Palindromic Repeats (CRISPR) is an adaptive immune system of bacteria. There are several types of CRISPR systems that vary in complexity. Out of the 5 systems discovered so far, type II consists of the fewest parts. This is why it has been chosen as the instrument of choice and what we’ll focus on here.
The CRISPR region consists of multiple copies of repeated, roughly palindromic sequences of 30 bases separated by spacer regions of about 36 bases. These curious regions have been found in several bacteria, which is why it caught the eye of scientists. If such a structure is so widely used, it must surely serve an important function!
The breakthrough moment was when scientists realised that the spacers are in fact copies of parts of the DNA of viruses that attack the bacteria. CRISPR saves parts of dangerous DNA so that it knows what to defend against the next time around. It serves as a memory of known aggressor DNA.
Cas
Knowing what to defend against can only be the first step. If CRISPR really was an immune system, there would need to be another part that would eventually disable the viruses. There must be an enzyme that would scan DNA in the cell and match it agains the list of aggressor DNA. If a match is found, the DNA should be destroyed.
There are enzymes that are CRISPR-associated (Cas) that play this role. Cas are nucleases, a type of enzyme that can cut DNA. The genes that encode for them are always close to the CRIPSR region. The most widely used type of Cas is Cas9. It’s part of the type II system that has the advantage of only requiring one Cas to work. The recently discovered Cpf1 enzyme might replace Cas9 as the standard of choice in gene editing, though.
![Cas9 Structure by Ben.lafrance [CC BY-SA 4.0 ] via Wikimedia Commons](https://res.cloudinary.com/dxrg6ryex/image/upload/v1566718833/Cas9_Apo_Structure_xxnvgb.png)
Bacterial defence process
So how does CRIPSR work in detail?
- The spacer DNA is copied into RNA. This RNA is used to guide Cas. That’s why it’s referred to as gRNA. (In nature there are two types of RNA that guide Cas, but scientists were able to combine them into one gRNA.)
- The gRNA binds to the Cas enzyme. This changes Cas into a state where it is active and can bind to DNA.
- The Cas/gRNA complex sift through the cell.
- If it encounters DNA, it binds to a short DNA sequence called PAM. It then unzips the DNA and proceeds to match DNA agains gRNA.
- If it finds a matching sequence, it cuts through both strands of the DNA forming a blunt end and disabling the virus.
There’s a really well-animated MIT video showing the whole process.
And a more in depth one by Jennifer Doudna, who discovered the gene engineering capabilities of CRISPR together with Emmanuelle Charpentier.
The nice thing about CRIPSR is that Cas is directed only by the gRNA. The only thing we’d need to program CRISPR is an artificial gRNA to guide it.
It’s really as simple as it sounds. Genius!
While being able to cut virus DNA is quite nice, CRISPR gained so much attention because scientists have shown that it can be used on a wide variety of cell types and species. CRISPR has been used to edit genomes of plants, fish, bacteria, pigs and humans. Not only that, but it even works inside the nucleus of a living cell that contains the DNA.
CRISPR advantages
CRISPR has many advantages over previously used gene editing techniques.
One advantage of the CRISPR is that Cas only matches “precisely”. The DNA is only cut when there’s an exact match between gRNA and the target DNA.
Another advantage is that the sequence used for targeting Cas9 can have a length of up to 20 nucleotides. This is relatively long. The longer the target the more precisely can we target the Cas9 to the specific region of the more than 3b base pairs in human DNA.
Moreover, several Cas9 can work simultaneously in one cell. By adding several gRNA we can edit DNA in different places in parallel. This allows us to study more complex DNA interactions that realistically match the processes going on in the cell.
Another advantage of CRISPR is that it’s really cheap to use. Cas9 is a standard component that doesn’t need to be adjusted per application. It can be bought off the shelf. The only thing that needs to be customised is the gRNA, which is relatively cheap, too. The total cost can be as little as $30.
Cutting? Don’t run with scissors!
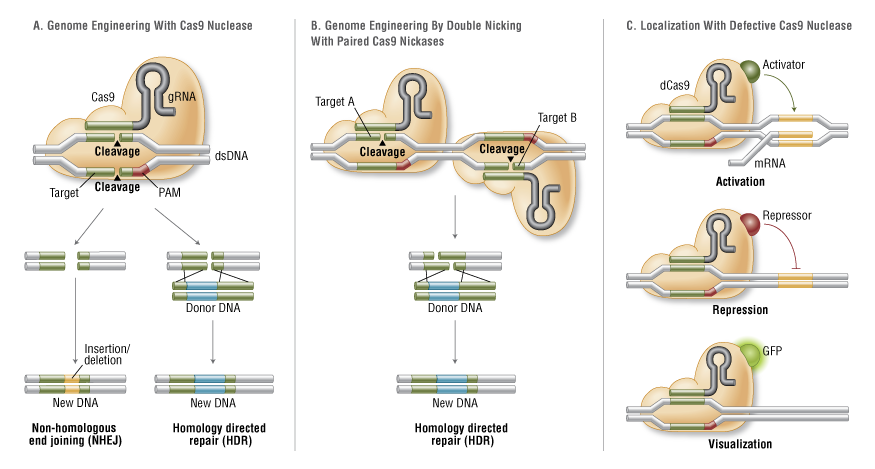
Using CRIPSR to cut DNA might not even be its most useful application. One reason why cutting is not always a good option, is that CRISPR produces double-stranded blunt ends.
Repairing these DNA breaks is error prone and can introduce unintended mutations. So scientist realised that the real advantage of CRISPR is it’s easy to program and precise targeting mechanism.
Having said that, CRISPRs have been developed that can only cut one DNA strand, which can increase specificity and mitigate the downsides of the repairs of double-stranded breaks.
Several different applications have beed developed for CRIPSR. A particularly exciting creation was that of a dead version of Cas9 - dCas.
While this version of the enzyme cannot cut DNA anymore, it can be used as a transport mechanism to target certain DNA sequences. The most important use case is the regulation of gene expression. We now can control genes, by activating or repressing them. Moreover, this platform can e.g. transport fluorescent proteins to a certain DNA region allowing us to observe it better.
Why it is important
CRIPSR is a breakthrough technology in many ways. Its versatility and easiness of use take gene editing to new levels. It provides us with the tools to edit genes inside the cell or other contexts in any organism.
(It’s actually so easy to use and took off so fast, that Dr. Doudna has called for a stop on using it until the scientific community has agreed on clear bioethical guidelines.)
There are already 2.000 gene therapies in development aiming at things like resistance to HIV, curing blindness or changing pig organs so that they can be transplanted into humans. The first gene therapy, Glybera, was already approved in Europe. With improvements such as CRISPR, the pace will only accelerate. The ability to edit several genes simultaneously is closer to how genes actually work. It allows us to write complex gene programs.
We can use it to not only to cut genes, but also to control and observe them.
This allows applications in many contexts far outside the widely reported ones in medicine. And that is exactly why it’s so exciting.